Genomic opportunities, part I: seeking the secrets of life
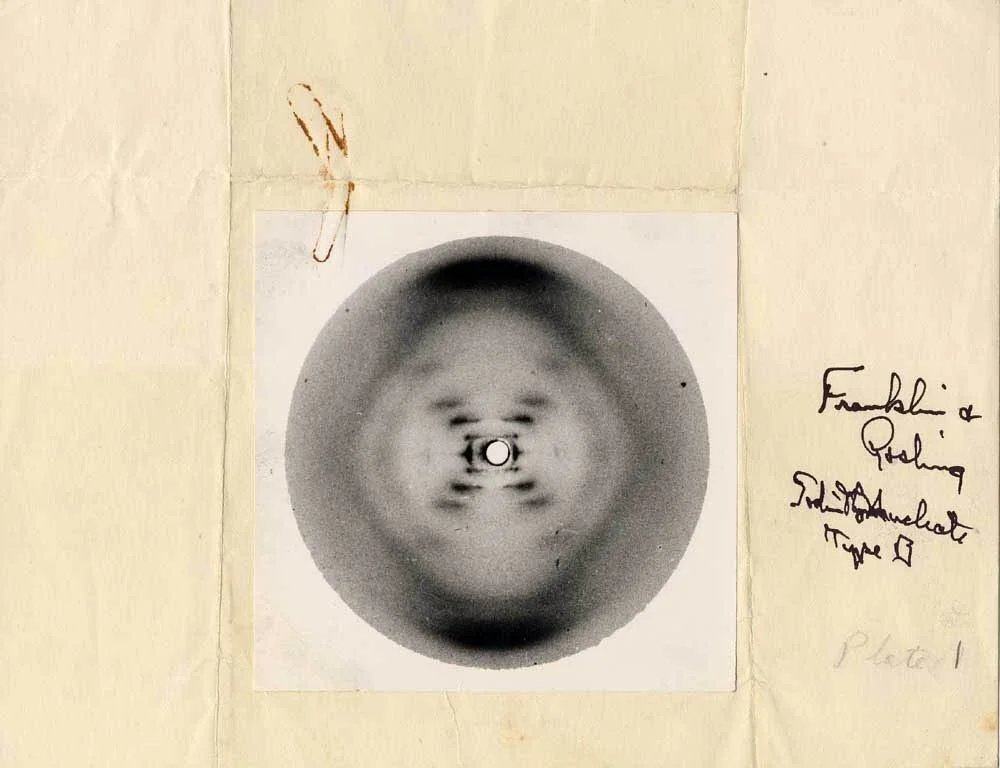
"We have discovered the secret of life!" Ever since Francis Crick burst into The Eagle pub one February afternoon in 1953, the genetic code has seemed to promise riches. Commercialisation of gene sequencing technologies and the development of computational genomics, predicated on the realisation that evolutionary molecular biology can be conceived as an information science, enabled the Human Genome Project, heralded at its culmination in 2000 as a full revelation of "the Book of Life" - that biblical metaphor originary to the scientific revolution itself. But neither claim was quite true: Life kept on keeping secrets.
"If there are many secrets left in the world, there are probably many world-changing companies yet to be started." At Mosaic, our job is to invest in these companies. In this and subsequent posts, we will explore the opportunities for building them out of genomics. What are the next companies discovering life's secrets?
One neat secret we used to talk about was "the gene for", perhaps for schizophrenia, cancer, height or musical ability. We know better now. Only a small number of diseases or positive traits are deterministically caused by single specific variants (e.g. haemophilia; lactase persistence). Nonetheless, the identification of these monogenic phenotypes has had important implications for diagnostics. Would-be parents from groups at risk for genetic disease are now generally screened, leading to some huge preventions achieved. But post-conception, commercially available non-invasive prenatal testing (NIPT), useful for de novo abnormalities in the large-scale structure of chromosomes (e.g. Down's Syndrome), is unable reliably to screen for monogenic conditions beyond paternally inherited autosomal dominant conditions, since it is predicated on DNA fragments circulating in the placenta. In addition, a large fraction of monogenic diseases are de novo mutations, and so would require a new kind of test to detect. Companies like Counsyl, Natera and Harmony (Roche) have been very successful (Natera is at ~$300m in annual revenues) selling diagnostics which are accurate for large structural abnormalities, but not comprehensive or performant in covering subchromosomal defects and, at best, very limited at the genetic level (in the UK, the largest private provider of NIPT recently stopped offering monogenic and subchromosomal screening, after attention was drawn to their low analytic performance). Despite a rapidly growing market, now over two million tests annually, current technology still cannot answer most of the questions a parent would want it to answer.
Therapies for these single gene disorders are even less developed. It was only in 2018 that the molecular basis of cystic fibrosis was actually understood (using single-cell methods); the long tail of still rarer genetic diseases is far less researched. (Companies like Healx are rising to meet the need for therapies for these, hoping to use AI to repurpose existing drugs hypothesis-free.) Monogenic diseases, in theory, admit of full cures via gene therapies, an area potentially on the cusp of a renaissance, with over 350 therapies in clinical development, and more than 30 at Phase III. A cure for haemophilia A could be approved by the middle of this year. We might also study rare genetic abilities, in particular predispositions for extraordinarily good health, as a means to discover biological mechanisms which might be harnessed for therapeutic benefit. (This is the stated goal of the recently unveiled Variant Bio.)
But if we zoom out, all this is just a tiny piece of the puzzle with which genomics has presented us. The overwhelming majority of phenotypic traits, positive and negative, are better considered polygenic (controlled by more than one gene), and subject to complex regulation as a function of cell type and changing environment. The famously faster-than-Moore's-Law decline in the cost of whole genome sequencing since 2007 (now itself declining due to Illumina's pricing power) has enabled the creation of the large population datasets which underlie the emergence of polygenic probabilistic risk scores today.

Genomics England has just announced an aspiration to sequence 5 million genomes in the UK over the next five years. The eventual application of GWAS-derived measures in clinical and potentially national screening contexts would be an important next step in the interventional authority of genomic screening, requiring new and careful communication: a high score means something categorically distinct to, say, a four base pair insertion in exon 11 (1278insTATC) (the most common allele for Tay-Sachs disease). This commercial trend arrives at a time when we have recently been reminded of an idea first described long before the molecular basis of the gene was known, in 1918 by Ronald Fisher. In the "omnigenic" model, every single base pair is causally involved in any complex trait.
So every gene is involved; but at least that's a finite number, and one that's computationally tractable (3 billion base pairs). Or, as psychiatrist Scott Alexander put it:
The "secret" of genetics was that there was no "secret". You just had to drop the optimistic assumption that there was any shortcut other than measuring all three billion different things, and get busy doing the measuring. The field was maximally perverse, but with enough advances in sequencing and computing, even the maximum possible level of perversity turned out to be within the limits of modern computing.
Unfortunately, this actually wasn't the maximum possible level of perversity biology could achieve: not by a long shot. The argument for biological causation being a (massively) bounded search problem would obtain if Crick's "Central Dogma" of monodirectional information flow was a reliable axiom. Unfortunately, there is, in fact, no such privileged level of causality in biological systems; no central conductor to the emergent "music of life". Nuclear DNA is constantly being regulated in new, fractally intricate ways we are still discovering each day, in response to the world around and inside the organism.
The reason we know this is because of an efflorescence of new tools for measuring biology in the past 15 years. Our ability to detect what is going on inside living systems has improved dramatically across two dimensions: new signals (RNA, proteomics, methylation, chromatin spatial organization etc.), and new resolutions (the single-cell sequencing wave; super-resolution microscopy). The ENCODE project which revealed the "dark matter" of the non-protein-coding genome in 2012, prompting, for many, a Copernican re-evaluation of what the genome is, would not have been possible without the combination and integration of of many of these technologies, (RNA-seq, CAGE, RNA-PET, ChIP-seq, DNase-seq, FAIRE-seq, MNase-seq etc). We believe these advances combined are, if anything, more significant than the staggering improvements in efficiency and cost of whole-genome sequencing; they have already redefined our understanding of biology, with tremendous implications for future applications, many still unexplored. The recent IPO and subsequent performance of 10x Genomics (shares are up nearly 80% from $5B since September last year) indicates the scale of possible outcomes in this infrastructure category; we think there are many other companies like this to be built.
As it scales both deeper and wider, genomics may soon be the biggest source of data on the planet (bigger than astronomy, Twitter and YouTube). But if living in the Information Age has taught us anything, it's that there is no necessary relation between more data and more insight. The success of 10x is in no small part due to its consistent attention to software as well as wetware. As the number of different signals being measured continues to grow, as experiments become exponentially multiplexed, we urgently need better tools to manage and integrate these vast pipelines of molecular data, together with phenotypic correlates, into rich (perhaps even "holistic") models which can handle the polyphonic texture of biological information processing, while at the same time keeping guarantees of robustness, privacy and security. The rate of growth has been dizzying: up until a couple of years ago, the whole pharmaceutical industry hadn't generated more than 100,000 genomes. This meant that bioinformatics-based companies until recently had a tiny market into which to sell; as a result, hacked-together pipelines were "good enough", and current infrastructure tooling is woefully inadequate to the grandeur of the tasks ahead. "It's amazing we were able to do all this with the tools we have", an exhausted bioinformatician friend recently remarked. Good carpenters don't blame their tools, but we have entered a regime where without new ones, they won't even be able to lift their material onto the workbench.
In the next post, we'll talk about these problems in bioinformatics infrastructure. Which challenges are unique to genomics, and not a special case of other kinds of software infrastructure tooling? Where are data generators building, where are they buying? What are the greatest unmet needs in turning raw molecular data into productive insight? After that, we'll discuss new reading technologies, and the transformative applications that new molecular signals and resolutions enable. Especially since the invention of the CRISPR-Cas9 technique, the idea of gene editing has dominated science headlines: we'll investigate new writing technologies, and their implications for genomics moving from a purely analytical to an interventional paradigm. Could there be an Illumina of synthetic biology? The next wave of gene-based medicine will be built on top of these categories; we'll discuss the role startups will play in this market. In the interest of exemplary focus, we've so far focused on medicine, but clearly genomics and synthetic biology together have the potential to reshape many other industries from the nucleus up in the medium term. At Mosaic, we have met companies hoping to rethink farming, meat, cleaning products, sweeteners, sensors, fuels and fashion: we'll lay out some hypotheses which we'd love to help entrepreneurs test.
If the 20th century was the century of physics, the 21st century will be the century of biology. While combustion, electricity and nuclear power defined scientific advance in the last century, the new biology of genome research - which will provide the complete genetic blueprint of a species, including the human species - will define the next. (Venter and Cohen, 1997)
We've pointed out how, with hindsight, the idea of a "complete blueprint" feels - rather like an artificial general intelligence - something we are unlikely to discover for a long time, if ever, but Venter's view of this century feels, 20 years in, truer by the day. Contemporary science fiction doesn't have much time for positive visions, but in today's nascent technologies, we should be able to see the reflected shadows of a feasible futurity: an immensely richer world where we've, at least, eliminated the burden of disease, developed agriculture that can feed the planet without destroying it, and reconceived manufacturing, inspired by the elegance of nature's factories. There is much work to do, many difficult but possible goals to pursue. Please get in touch if you have secrets to tell.